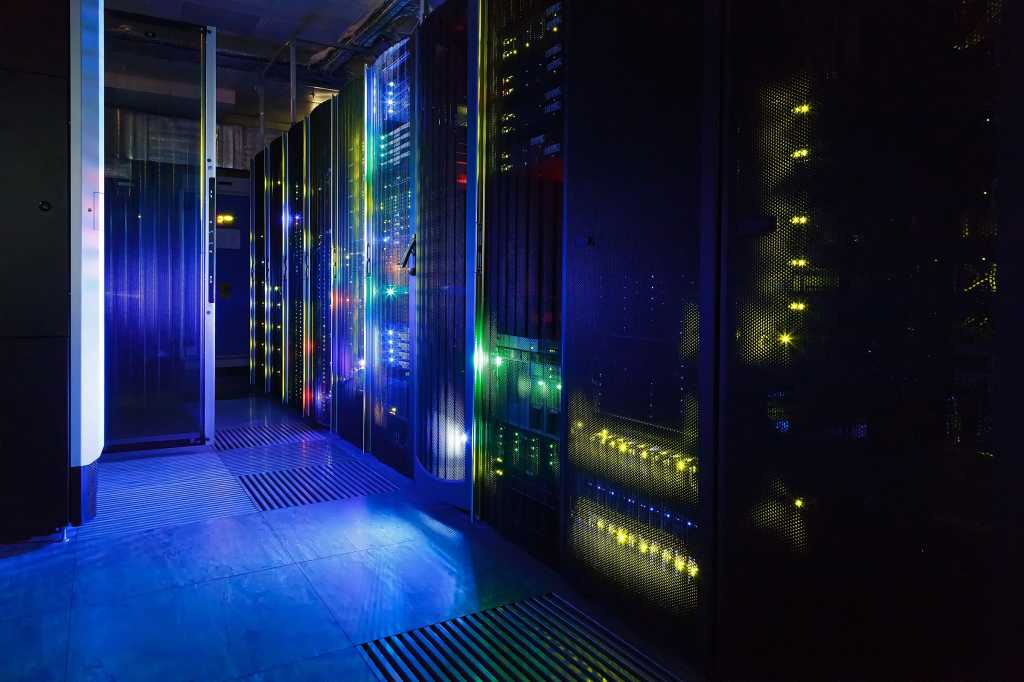
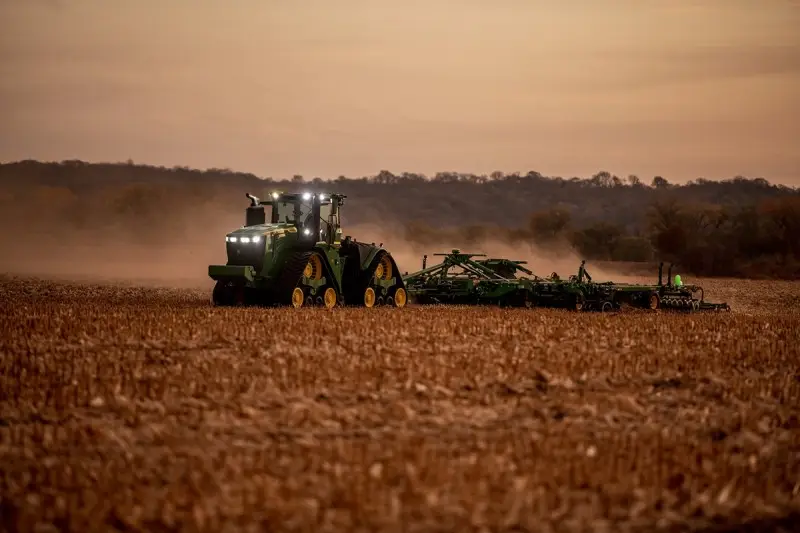
Microbes that gobble up or break down environmental toxins can clean up oil spills, waste sites, and contaminated watersheds. But until his faculty mentor asked him for help with a project he was working on with doctors at Boston Children’s Hospital in 2009, Eric Alm had not thought much about their role in a very different environment: the human digestive system.
David Schauer, a professor of biological engineering, was examining how microorganisms in the gut might be linked to inflammatory bowel disease (IBD), and he hoped advanced statistical analysis of the data he was collecting could make those connections clearer. Alm, who’d joined the civil and environmental engineering faculty in 2006 as a computational biologist studying environmental uses of microbes, had the statistical experience needed and could apply machine-learning tools to help. But for him, the project was supposed to be a brief detour.
In June of 2009, however, Schauer—just 48—died unexpectedly, only two weeks after falling ill. Alm, heartbroken, worked to help push his mentor’s project over the finish line. As that effort was underway, Neil Rasmussen ’76, SM ’80, a longtime member of the MIT Corporation and the philanthropist funding the project, asked for a tour of his lab. That encounter would change the course of Alm’s career.
At the end of the lab tour, Rasmussen, who has a family member with IBD, had a surprise: He asked Alm if he’d be willing to pivot to researching inflammatory bowel disease—and offered to fund his lab if he did so.
Alm was game. He began shifting the main focus of his research away from harnessing microbes for the environment and turned most of his attention to exploring how they could be applied to human health. Then Rasmussen decided he wanted to “do something really big,” as Alm puts it, and make Boston a hub for microbiome research. So in 2014, with a $25 million grant from the Neil and Anna Rasmussen Foundation, the Center for Microbiome Informatics and Therapeutics (CMIT) was launched with Alm and Ramnik Xavier, chief of gastroenterology at Massachusetts General Hospital, as its co-directors.

By teaming up with Alm and others, Rasmussen hoped to create a research hub where scientists, engineers, doctors, and next-generation trainees would collaborate across scientific disciplines. They would build the tools needed to support a new research field and translate cutting-edge research into clinic-ready interventions for patients suffering from a wide range of inflammatory and autoimmune conditions influenced by the gut, including not only IBD but diabetes and Alzheimer’s—and potentially autism, Parkinson’s disease, and depression as well.
In its first 10 years, CMIT has made remarkable progress.
When the center started, Alm says, it was still a relatively novel idea that the human microbiome—particularly the community of trillions of symbiotic microbes that reside in the gut—might play a key role in human health. Few serious research programs existed to study this idea.
“It was really this undiscovered territory,” he recalls. “[In] a lot of diseases where there seemed to be things that we couldn’t explain, a lot of people thought maybe the microbiome plays a role either directly or indirectly.”
It has since become increasingly clear that the microbiome has a far greater impact on human health and development than previously thought. We now know that the human gut—often defined as the series of food-processing organs that make up the gastrointestinal tract—is home to untold trillions of microorganisms, each one a living laboratory capable of ingesting nutrients, sugars, and organic materials, digesting them, and releasing various kinds of organic outputs. And the metabolic outputs of these gut-dwelling microbes are similar to those of the liver, Alm says. In fact, the gut microbiome can essentially mirror some of the liver’s functions, helping the body metabolize carbohydrates, proteins, and fats by breaking down complex compounds into simpler molecules it can process more easily. But the gut’s outputs can change in either helpful or harmful ways if different microbes establish themselves within it.
“I would love to have bacteria that live on my face and release sunscreen in response to light. Why can’t I have that?”
Tami Lieberman
“Our exquisite immune defenses evolved in response to the microbiome and continue to adapt during our lifetime,” Rasmussen says. “I believe that advancing the basic science of human interactions with the microbiome is central to understanding and curing chronic immune-related diseases.”
By now, researchers affiliated with the center have published some 200 scientific papers, and it has found ways to advance microbiome research far beyond its walls. It funds a team at the Broad Institute (where Alm is now an Institute Member) that does assays and gene sequencing for scientists doing such research. Meanwhile, it has established one of the world’s most comprehensive microbiome “strain libraries,” facilitating studies around the globe.
To create this library—which includes strains in both the Broad Institute–OpenBiome Microbiome Library and the Global Microbiome Conservancy’s Biobank—researchers have isolated more than 15,000 distinct strains of microbes that are found in the human gut. The library can serve as a reference for those hoping to gain information on microbes they have isolated on their own, but researchers can also use it if they need samples of specific strains to study. To supplement the strain library, CMIT-affiliated researchers have traveled to many corners of the globe to collect stool samples from far-flung indigenous populations, an effort that continues to this day through the Global Microbiome Conservancy.
“We’re trying to build a critical mass and give folks working in different labs a central place where they can communicate and collaborate,” says Alm. “We also want to help them have access to doctors who might have samples they can use, or doctors who might have problems that need an engineering solution.”
The clinical applications produced by CMIT have already affected the lives of tens of thousands of patients. One of the most significant began making an impact even before the center’s official launch.
For decades, hospitals had been grappling with the deadly toll of bacterial infections caused by Clostridioides difficile (C. diff), a hardy, opportunistic bacterium that can colonize the gut of vulnerable patients, often after heavy doses of antibiotics wipe out beneficial microbes that usually keep C. diff at bay. The condition, which causes watery diarrhea, abdominal pain, fever, and nausea, can be resistant to conventional treatments. It kills roughly 30,000 Americans every year.
By 2003, researchers had discovered that transplanting stool from a healthy donor into the colon of a sick patient could restore the healthy microbes and solve the problem. But even a decade later, there was no standardized treatment or protocol—relatives were often asked to bring in their own stool in ice cream containers. In 2013, Mark Smith, PhD ’14, then a graduate student in Alm’s lab, cofounded the nonprofit OpenBiome, the nation’s first human stool bank. OpenBiome developed rigorous methods to screen donors (people joke that it’s harder to get approved than to get into MIT or Harvard) and standardized the procedures for sample processing and storage. Over the years, the nonprofit has worked with some 1,300 health-care facilities and research institutions and facilitated the treatment of more than 70,000 patients—work that OpenBiome says helped set the stage for the US Food and Drug Administration to approve the first microbiome-based therapeutic for recurrent C. diff infections.
Today, CMIT’s flagship effort is a 100-patient clinical trial that it launched to study IBD, using a wide array of technologies to monitor two cohorts of patients—one in the US and the other in the Netherlands—over the course of a year. People with Crohn’s disease and ulcerative colitis typically experience periods of full or partial remission, but they currently have no way to predict when they will relapse. So researchers are tracking weekly changes in each patient’s microbiome and other biological indicators while amassing continuous physiological data from Fitbits and recording self-reported symptom scores along with other clinical data. The goal is to identify biomarkers and other indicators that might be used to predict flare-ups so that already approved therapies can be used more effectively.
Although data is still being collected, early analysis suggests that a patient’s gut microbiome begins to change six to eight weeks before flare symptoms appear, and a few weeks later, genetic analysis of epithelial cells in their stool samples starts to show signs of increased inflammation. The team is planning to host a hackathon this summer to help speed analysis of the mountain of disparate types of data being collected.
Meanwhile, the community of clinicians, engineers, and scientists CMIT has nurtured is undertaking projects that Alm could hardly have imagined when he first delved into research on the human microbiome.
Survivor: Microbe edition
Right below the photograph on the bio page of her Twitter/X account, Alyssa Haynes Mitchell has three emojis: a tiny laptop, a red and blue strand of DNA, and a smiling pile of poo. The digital hieroglyphics neatly sum up her area of focus as she pursues a doctorate in microbiology. A 2024 Neil and Anna Rasmussen fellow, Mitchell is attempting to understand precisely what it is that allows microbes to survive and thrive in the human gut.
Mitchell fell in love with the study of microbes as an undergrad at Boston University. First, her mind was blown after she read a paper by researchers who could create a facsimile of a patient’s intestinal cell population—a “gut on a chip”—and planned to culture a microbiome on it. She was fascinated by the idea that this might lead to personalized treatments for conditions like IBD. Then she cultured her first colony of a strain of the microbe Bacillus subtilis that had been genetically engineered to fluoresce.
“They form these really complex ridges, and the more you look at microscopy images, the more you realize that there’s patterns of collective behavior of bacterial biofilms that we just don’t understand,” she says. “They’re super beautiful, and it’s really quite amazing to look at.”
In 2023, Mitchell joined the lab of Tami Lieberman, an associate professor of civil and environmental engineering and a member of both CMIT and MIT’s Institute for Medical Engineering and Science.
Mitchell and others who study the microbiome think that “probiotics,” beneficial microbes that are applied to the skin or ingested in supplements or foods such as yogurt or kombucha, could have broad potential to help treat disease. But for reasons that still aren’t well understood, once probiotics are introduced into the gut, only a small percentage of them are able to survive and proliferate, a process known as engraftment. A probiotic with an engraftment rate of 30% (meaning it’s still detectable in 30% of subjects) six months after administration is considered good, says Mitchell. She and Lieberman, who also holds the title of Hermann L.F. von Helmholtz Professor, are studying the way individual strains of microbes evolve to survive in the microbiome—a key mystery that needs to be solved to engineer more effective, longer-lasting therapies.


Alyssa Haynes Mitchell, a PhD student pursuing a doctorate in microbiology, is working with Tami Lieberman, an assistant professor of civil and environmental engineering, to study how strains of microbes evolve to survive in the gut. Lieberman also studies how microbes survive and proliferate on the skin.
“Hopefully if we learn a little bit more about what drives evolution of the ones that stick around, we might be able to learn why some don’t,” she says.
Mitchell has been working with samples collected by a local biotech company developing biotherapeutics for the gut. Its probiotic products, which are used to treat recurrent C. diff infections, contain eight closely related microbial strains belonging to the order known as Clostridiales. The company gave one of its products to 56 human subjects and collected stool samples over time. Mitchell is using genetic sequencing techniques to track how three of the microbial species evolved in 21 of the subjects. Identifying person-specific differences and similarities might reveal insights about the host environment and could help explain why some types of mutations allow some microbes to survive and thrive. The project is still in its early phases, but Mitchell has a working hypothesis.
“The model that I have in my mind is that people have different [gut] environments, and microbes are either compatible with them or not,” she says. “And there’s a window in which, if you’re a microbe, you might be able to stick around but maybe not thrive. And then evolution kind of gets you there. You might not be very fit when you land there, but you’re close enough to hang around and get there. Whereas in other people, you’re totally incompatible with what’s already there, and the resident microbes beat you out.”
Her work is just one of many projects using new approaches developed by Lieberman, who worked as a postdoc in Alm’s lab before starting her own in 2018. As a graduate student at Harvard, Lieberman gained access to more than 100 frozen samples collected from the airways, blood, and chest tissue of 14 patients with cystic fibrosis, a genetic disease that causes mucus to build up in the lungs and creates conditions ripe for infections. The patients were among those who had developed bacterial infections during an outbreak in the 1990s.
Lieberman and her colleagues recognized a perfect opportunity to use genetic sequencing technologies to study the way the genome of the Burkholderia dolosa bacterium evolved when she cultured those samples. What was it that allowed B. dolosa to adapt and survive? Many of the surviving microbes, she discovered, had developed similar mutations independently in different patients, suggesting that at least some of these mutations helped them to thrive. The research indicated which genes were worthy of further study—and suggested that this approach holds promise for understanding what it takes for microbes to grow well in the human body.
Lieberman joined Alm’s lab in 2015, aiming to apply the same experimental paradigm and the statistical techniques she had developed to the emerging field of microbiome research. In her own lab, she has developed an approach to figuring out how the pressures of natural selection result in mutations that may help certain microbes to engraft. It involves studying colonies of bacteria that form on the human skin.
“The idea is to create a genetically engineered metabolite factory in the gut.”
Daniel Pascal
In the gut, Lieberman explains, hundreds of different species of microbes coexist and coevolve, forming a heterogeneous community whose members interact with one another in ways that are not fully understood. This creates a wide array of confounding variables that make it more difficult to identify why some engraft and others don’t. But on the skin, the metabolic environment is less complex, so fewer species of bacteria coexist. The smaller number of species makes it far easier to track the way the genomes of specific microbes change over time to facilitate survival, and the accessibility of the skin makes it easier to figure out how spatial structure and the presence of other microbes affect this process.
One discovery from Lieberman’s lab is that each pore is dominated by just one random strain of a single species. Her group hypothesizes that survival may depend on the geometry of the pore and the location of the microbes. For example, as these anaerobic microbes typically thrive at the hard-to-access bottom of the pore, where there is less oxygen, the first to manage to get there can crowd out new migrants.
“My vision, and really a vision for the microbiome field in general,” Lieberman says, is that one day therapeutic microbes could be added to the body to treat medical conditions. “These could be microbes that are naturally occurring, or they could be genetically engineered microbes that have some property we want,” she adds. “But how to actually do that is really challenging because we don’t understand the ecology of the system.” Most bacteria introduced into a person’s system, even those taken from another healthy human, will not persist in the new person’s body, she notes, unless you “first bomb it with antibiotics” to get rid of most of the microbes that are already there. “Why that is,” she adds, “is something we really don’t understand.”
If Lieberman can solve the puzzle, the possible applications are tantalizing.
“I would love to have bacteria that live on my face and release sunscreen in response to light,” she says. “Why can’t I have that? In the future, there’s no reason we can’t figure out how to do that in a safe and controlled manner. And it would be much more convenient than applying sunscreen every day.”
Harnessing light-sensitive, sunblock-producing microbes may sound like a distant fantasy. But it’s not beyond the realm of possibility. Other microbial products that sound straight out of a science fiction novel have already been invented in the lab.
Molecular assassin
When Daniel Pascal first landed in the lab of MIT synthetic biologist Christopher Voigt, he had no idea he’d be staying on to make bacteria with superpowers. He was a first-year PhD student rotating through various labs, with little inkling of the potential contained in the microbes that live inside us.
Pascal, a 2024 Neil and Anna Rasmussen fellow who is pursuing a doctorate in biological engineering, was originally paired with a graduate student doing a more materials-related synthetic biology project. But he came from a family of physicians and soon found himself speaking with other graduate students in the lab whose projects had to do with health.
He then learned that two of the lab’s postdocs, Arash Farhadi and Brandon Fields, were receiving funding under a program sponsored by the Defense Advanced Research Projects Agency (DARPA), the Pentagon’s R&D organization, to develop solutions for common traveler’s ailments that result from problems like disrupted sleep cycles and limited access to safe food and water. When they explained that they hoped to harness microbes in the human body, they had his attention.

“It’s amazing how these tiny little organisms have so much control and can wreak so much havoc,” he says.
Intrigued, Pascal wound up officially joining Voigt’s lab, where he is working to create microbes that can carry out a wide array of functions they would not perform in the natural world.
To do so, he is using a custom “landing pad” system developed in the lab. The system relies on synthetic biology to create a new region in the genome of a microbe that, using specific enzymes, can be filled with pieces of DNA designed to imbue the microbe with special new abilities.
After engineering the landing pad into samples of an existing probiotic, Pascal and his collaborators on a project funded by the US Air Force and DARPA were able to deliver DNA that allows the probiotic to essentially set up a specialized drug production facility within the gut. First it absorbs two common amino acids, arginine and glycine. Then it converts them into a precursor compound that the body transforms into creatine, which can facilitate the production of muscle tissue from exercise and may help with memory.
Pascal explains that creatine is often taken as an over-the-counter supplement by people doing weight training and other athletes who want to improve their fitness. “But creatine has been shown to improve performance in fatigued humans,” he says. “So the motivation for this project was the idea that Air Force pilots that are traveling all over the world are jet-lagged, are working crazy hours and shifts.” What if, the researchers wondered, those pilots “could take a supplement that would improve some of their responsiveness, athletic accuracy, intelligence, and reasoning?”
A typical oral supplement delivers a spike of creatine in the bloodstream that largely dissipates relatively quickly. More useful to the pilots would be a probiotic engineered to produce a consistent amount of the creatine precursor that could be turned into creatine as needed.
CMIT is also funding Pascal’s project using the landing pad system to get microbes to produce substances that target specific pathogens without disrupting the entire microbiome. Although Pascal cannot yet reveal any details about these molecular-level assassins, he notes that other researchers in the Voigt lab have recently used the landing pad system to redesign the Escherichia coli Nissle (EcN) microbe, which had previously been engineered to produce such things as antibiotics, enzymes that break down toxins, and chemotherapy drugs to fight cancer. The lab’s work made it possible to improve the efficacy of a treatment for phenylketonuria and perhaps of other EcN therapeutics as well.
The lab has, in short, been able to get microbe strains (one of which he says is a commercially available probiotic that in some countries you can buy over the counter) to do some very useful things. “They’ve figured out a way to take this mundane thing and give it these extraordinary capabilities,” he says. “The idea is to create a genetically engineered metabolite factory in the gut.”
Tackling childhood obesity
Understanding the microbiome may also lead to new therapies for one of the greatest public health challenges currently facing the US: rising rates of obesity.
Jason Zhang, a pediatric gastroenterologist at Boston Children’s Hospital, has received a CMIT clinical fellowship to study how gut bacteria may be linked to childhood obesity and diabetes. As a visiting scientist in Alm’s lab, he is using AI to predict people’s loss of control over what or how much they eat. His working hypothesis is that microbial metabolites are interacting with endocrine cells in the lining of the gut. Those endocrine cells in turn secrete hormones that travel to the brain and stimulate or suppress hunger.
“We believe that the microbiome plays a role in how we make choices around food,” he says. “The microbiome can send metabolites into the bloodstream that will maybe cross the blood-brain barrier. And there may be a direct connection. There is some evidence of that. But more likely they’re going to be interacting with cells in the epithelial layer in the gut.”

Zhang has sequenced the microbes found in the stool of subjects who have exhibited “loss-of-control eating” and developed a machine-learning algorithm that can predict it in other patients on the basis of their stool samples. He and his colleagues have begun to home in on a specific microbe that appears to be deficient in kids who experience this eating pattern.
The researchers have discovered that this particular microbe appears to respond to food in the gut by creating compounds that stimulate enteroendocrine cells to release a series of hormones signaling satiety to the brain—among them GLP-1, the hormone whose signal is turned up by weight-loss drugs like Ozempic. Zhang has already begun experimenting with therapies that artificially introduce the microbe into mice to treat obesity, diabetes, and food addiction.
“As with any single mechanism that treats a really complex disease, I would say it’s likely to make a difference,” he says. “But is it the silver bullet? Probably not.” Still, Zhang isn’t ruling it out: “We don’t know yet. That’s the ongoing work.”
All these projects provide a taste of what’s to come. For more than a decade, CMIT has played a key role in building the fundamental infrastructure needed to develop the new field.But with as many as 100 trillion bacterial cells in the human microbiome, the efforts to explore it have only just begun.